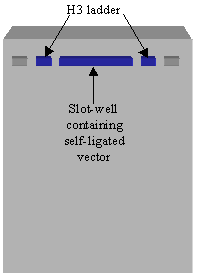

(a) Prepare a "slot-well" by joining several comb-made wells together. Seal the bottom of the slot-well with a thin layer of melted agarose. Place the vector DNA (with blue juice) in the slot-well. Place the H3 ladder in wells flanking the slot well.
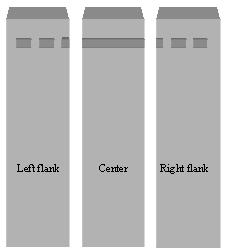
(b) After electrophoresis, divide the gel into three pieces as shown. The two flanking pieces should each contain a small part of the slot-well as well as a DNA ladder. Stain the flanking pieces with ethidium bromide. Do not stain the center piece (the gel piece containing most of the vector DNA).
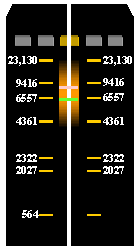
(c) Align the stained flanking gel pieces and examine on a UV light box. There should be a distinct band at approximately 7.5 kb marking the dephosphorylated vector. Using a scalpel, make incisions in the flanking pieces above and below this band (pink and green lines, respectively). Linear concatemers may be visible at (7.5 x 2 =) 15 kb and (7.5 x 3 =) 22.5 kb. Additionally, circular DNA molecules may be appear at molecular weights > 7.5 kb.
(d) Using the incisions in the flanking pieces as guides and a ruler as a "straight edge", cut the 7.5 kb band containing dephosphorylated vector out of the unstained center piece of the gel. DNA is isolated from the gel band containing dephosphorylated vector using a combination of freezing and centrifugation (see METHODS).
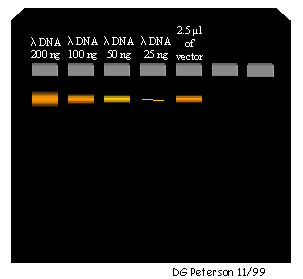
FIGURE 3.2 - Diagram of a mini-gel used to determine the DNA concentration of the dephosphorylated vector DNA. Four different concentrations of uncut lambda DNA are run in the first four lanes while 2.5 µl of the vector is run in lane 5. In this example, 2.5 µl of purified vector stains with an intensity roughly equal to that of the 100 ng of 1X uncut lambda DNA. Because 2.5 µl of vector DNA was placed in lane 5, the concentration of the undiluted DNA sample is about (100 ng/2.5 µl =) 40 ng/µl.