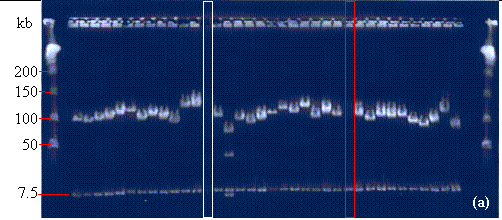

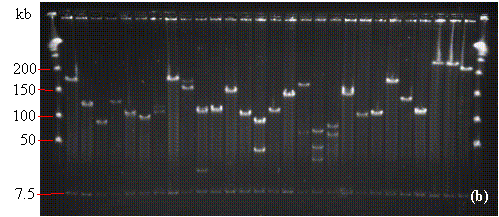
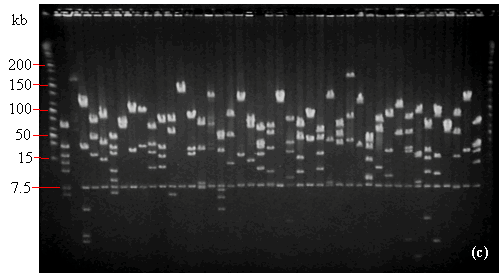
FIGURE 14.1
- NotI digests of minipreps from randomly-selected white colonies of (a) Gossypium raimondii, (b) soybean (Glycine max 'A3244'), and (c) maize (Zea mays 'B73') libraries. In (a) and (b) the PFGE lambda ladder is visible at the far-right and far-left sides of the photograph. In (c), the PFGE Midrange I ladder is in the first and last lanes. In each photo, note the row of vector bands at 7.5 kb. For the dicots G. raimondii (a) and soybean (b), most lanes contain only one or two non-vector bands indicating relatively few genomic NotI sites. As shown in (c), NotI restriction sites are considerably more common in monocot DNA. In (a), one of the lanes contains no visible DNA suggesting that the miniprep was lost (white rectangle), and one "false positive" (red rectangle) is visible. No false positives or empty lanes are seen in (b) and (c). The average insert sizes for the G. raimondii, soybean, and maize libraries are 120, 140, and 140 kb respectively (see TABLE 1.1).